Tutorial on LAI Visualization¶
import os
import sys
import logging
logging.basicConfig(stream=sys.stdout, level=logging.INFO)
dir = os.path.abspath('../')
if not dir in sys.path: sys.path.append(dir)
from snputils.ancestry.io.local.read import MSPReader
from snputils.visualization.lai import plot_lai
INFO:numexpr.utils:Note: detected 72 virtual cores but NumExpr set to maximum of 64, check "NUMEXPR_MAX_THREADS" environment variable.
INFO:numexpr.utils:Note: NumExpr detected 72 cores but "NUMEXPR_MAX_THREADS" not set, so enforcing safe limit of 16.
INFO:numexpr.utils:NumExpr defaulting to 16 threads.
1. Load Data¶
Load the Local Ancestry Inference (LAI) data into a LocalAncestryObject. This data will be used as input to plot_lai.
# Specify the path to the LAI data file
filename = '../data/easComp_6_samples_chr1.msp'
# Load LAI data using MSPReader, which returns a LocalAncestryObject
laiobj = MSPReader(filename).read()
INFO:snputils.ancestry.io.local.read.msp:Reading '../data/easComp_6_samples_chr1.msp'...
laiobj.haplotypes
['GA000856_GA000856.0',
'GA000856_GA000856.1',
'GA000857_GA000857.0',
'GA000857_GA000857.1',
'GA000858_GA000858.0',
'GA000858_GA000858.1']
# Construct haplotypes and LAI data
samples = list(range(10))
haplotypes = [f"{sample}_{sample}.0" for sample in samples] + [f"{sample}_{sample}.1" for sample in samples]
2. Define Ancestry Colors¶
Create a color mapping for each ancestry. This dictionary assigns a specific color to each ancestry label, which will be used in the visualization.
# Define colors for each ancestry type to use in the plot
colors = {
'Africa': "#87096c",
'Americas': "#9DADA5",
'Europe': "#EDD87E",
'SouthAsia': 'orange',
'EastAsia': 'green'
}
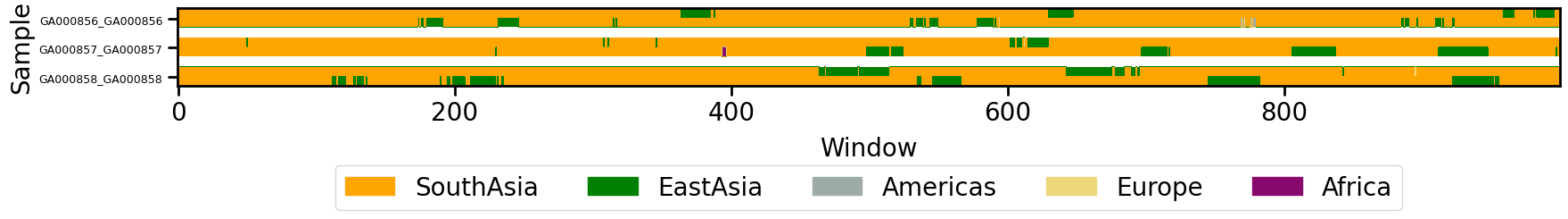
3. Visualize the LAI Data¶
Use the plot_lai function to create a visual representation of LAI data, with options for sorting by predominant ancestry, scaling, and configuring the display.
Explanation of Parameters:
colors: A dictionary mapping ancestry labels to specific colors, allowing clear differentiation between ancestries in the plot.sort: When True, samples are displayed in order of the most predominant ancestry, enhancing interpretability.figsize: Controls the size of the plot, which is important for visibility, especially with a large number of samples.legend: When True, a legend appears showing the color for each ancestry.scale: Determines how many times each row is repeated, which can enhance vertical visibility for individual samples.
plot_lai(laiobj, colors=colors, sort=True, figsize=(20, 100), legend=True,
title=None, fontsize=None, scale=7)